Synthetic Biology strives to develop artificial biological systems for applications in biotechnology and medicine. Successful engineering of novel cellular pathways requires genetic devices that optimize the expression of individual pathway modules.
We are interested in developing novel RNA-based devices, which can be controlled by exogenous ligands. In order to achieve highly specific ligand recognition and orthogonality between parts, we use RNA aptamers as sensor units. RNA aptamers are in vitro selected RNA molecules that, reminiscent of antibodies, recognize a particular ligand with high specificity and affinity.
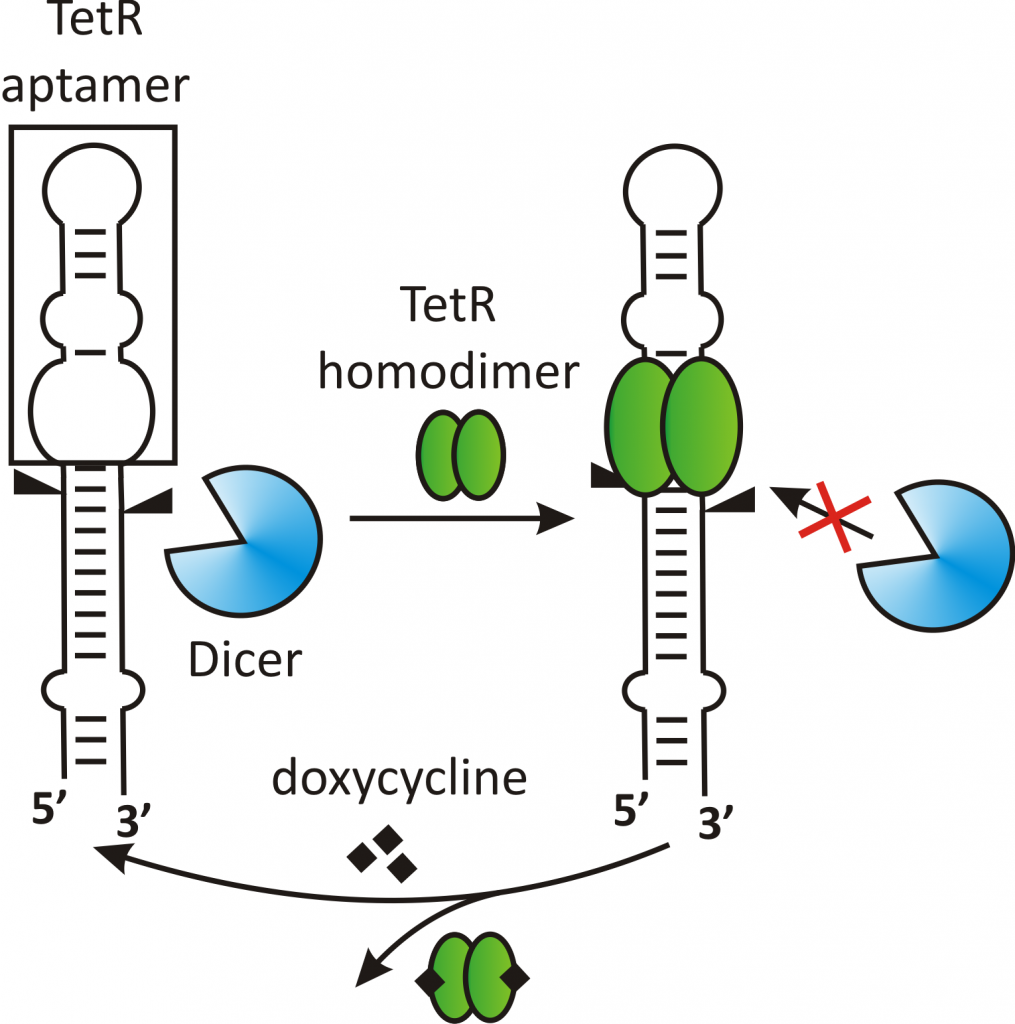
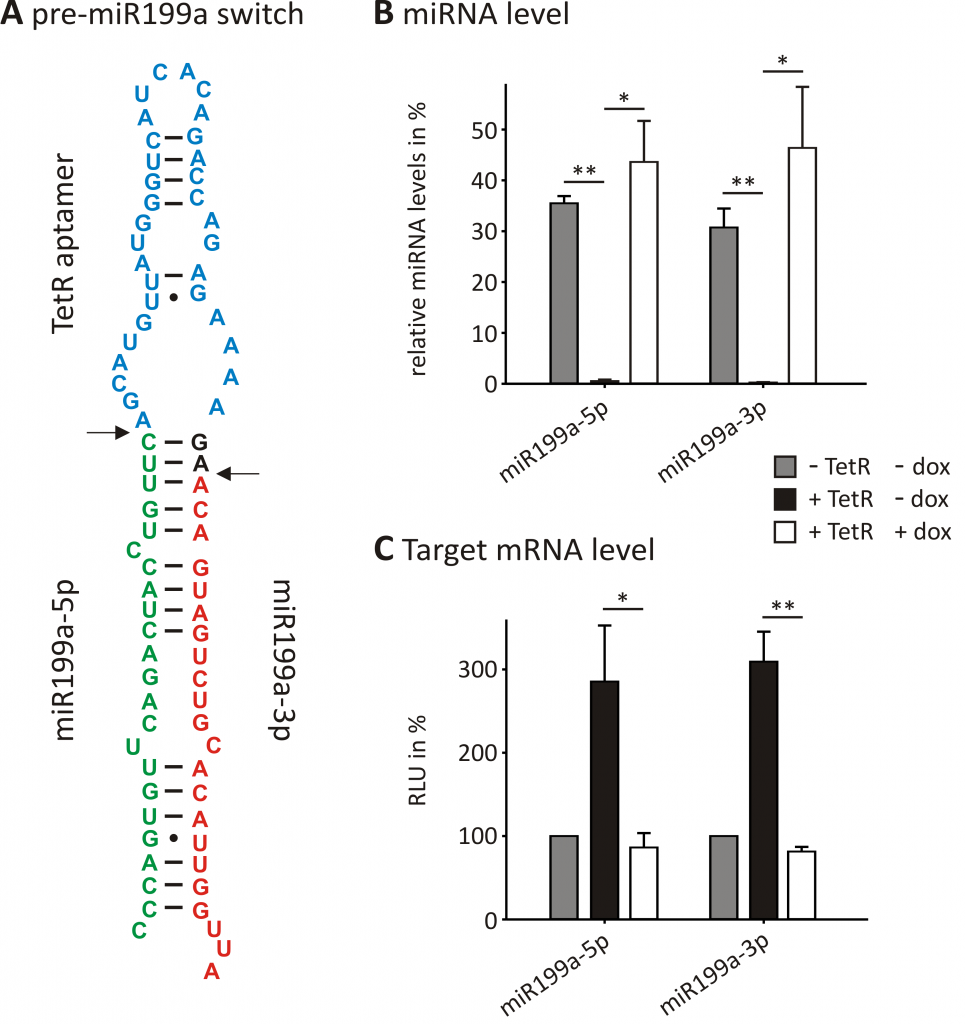
For example, we used an aptamer recognized by the bacterial repressor TetR to control the maturation of miRNAs (Fig. 1). miRNAs are small, non-coding RNAs that suppress gene expression by binding directly to the 3’UTRs of mRNAs. The mature miRNAs are processed from a long primary RNA transcript in a two-step process. The functionalization of such a transcript with the TetR aptamer enabled us to reversibly control the second processing step. This enables us to control the miRNA and, thereby, the target mRNA level within cells (Fig. 2). In future research, we intend to integrate our newly identified post-transcriptional elements from FOCUS 1-3 into genetic circuits and thus expand the toolbox of Synthetic Biology by adding novel, well characterized parts.